Masashi Maekawa
Development of a protein knockdown method by bivalent DNA aptamers
 |
Masashi Maekawa, PhDDivision of Physiological Chemistry and Metabolism, Graduate School of Pharmaceutical Sciences, Keio University |
|---|
Research summary
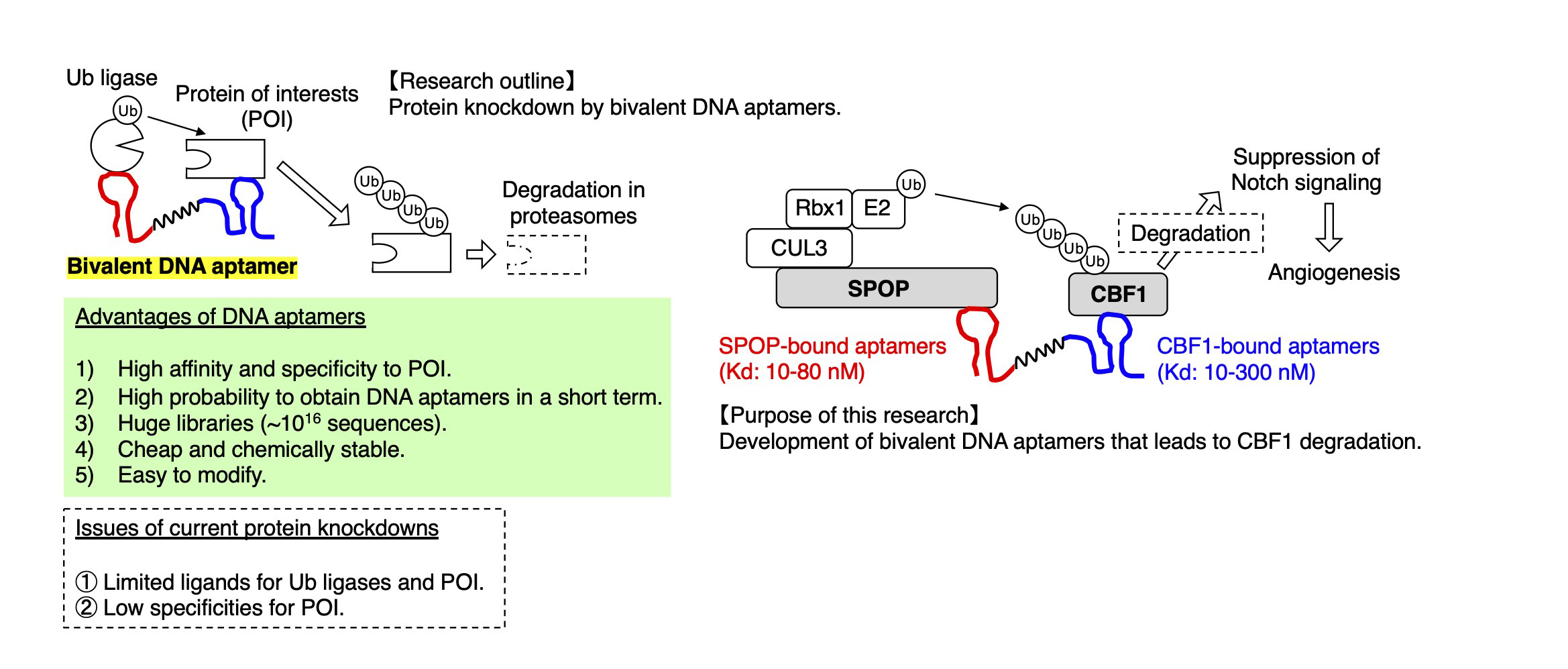
Protein knockdown is a newly-emerging and attractive method to degrade proteins of interests (POI). The method makes use of ubiquitin-proteasome pathways or autophagy pathways to achieve the degradation of POI. Although protein knockdown has various advantages from both clinical standpoints and basic research-standpoint, the method still has a couple of issues. Especially, the limited numbers of ligands for both ubiquitin ligases and POI pose a major challenge.
DNA aptamer is a single-stranded deoxyribonucleic acid (70-100 mers) that specifically binds to target molecules such as POI. The DNA aptamers have various advantages owing to its selection system, called systematic evolution of ligands by exponential enrichment (SELEX). In SELEX, the initial library pool contains 1014–1016 sequences, and thus, we are able to obtain DNA aptamers that specifically and directly bind to POI with high affinity.
In this research proposal, we hypothesized that bivalent DNA aptamers, which bridge two different DNA aptamers for ubiquitin ligases and POI, could keep ubiquitin ligases and POI closer leading to polyubiquitination of POI followed by its proteasomal degradation. To prove this concept, we used CBF1-bound DNA aptamers and SPOP-bound DNA aptamers as ligands for POI and a ubiquitin ligase, respectively. CBF1 is a transcription factor of Notch signaling that negatively regulates angiogenesis. SPOP is a substrate recognition receptor for a cullin-3/RING ubiquitin ligase complex. We develop bivalent DNA aptamers that crosslink CBF1-bound DNA aptamers and SPOP-bound DNA aptamers. As readouts, we examine SPOP-dependent polyubiquitination of CBF1 in vitro and CBF1 degradation in endothelial cells by making use of lipid nanoparticles. We aim to establish a new protein knockdown method by bivalent DNA aptamers.
Publications
- Nishiyama K, *Maekawa M (co-first), Nakagita T, Nakayama J, Kiyoi T, Chosei M, Murakami A, Kamei Y, Takeda H, Takada Y, *Higashiyama S.
CNKSR1 serves as a scaffold to activate an EGFR phosphatase via exclusive interaction with RhoB-GTP.
Life Sci. Alliance.4, e202101095 (2021)
PMID: 34187934 - Sanada S, *Maekawa M (co-first), Tate S, Nakaoka H, Fujisawa Y, Sayama K, *Higashiyama S.
SPOP is essential for DNA replication licensing through maintaining translation of CDT1 and CDC6 in HaCaT cells.
Biochem. Biophys. Res. Commun. 651, 30-38 (2023)
PMID: 36791496
Former Publications
- Maekawa M, Terasaka S, Mochizuki Y, Kawai K, Ikeda Y, Araki N, Skolnik EY, *Taguchi T, *Arai H.
Sequential breakdown of 3-phospholylated phosphoinositides is essential for the completion of macropinocytosis.
Proc. Natl. Acad. Sci. USA. 111, E978-E987 (2014)
PMID: 24591580 - Maekawa M, *Fairn GD.
Complementary probes reveal that phosphatidylserine is required for proper transbilayer distribution of cholesterol.
J. Cell Sci. 128, 1422-1433 (2015)
PMID: 25663704 - Murakami A, *Maekawa M (co-first), Kawai K, Nakayama J, Araki N, Semba K, Taguchi T, Kamei Y, Takada Y, *Higashiyama S.
Cullin-3/KCTD10 E3 complex is essential for Rac1 activation through RhoB degradation in human epidermal growth factor receptor 2-positive breast cancer cells.
Cancer Sci. 110, 650-661 (2019)
PMID: 30515933 - Watanabe R, *Maekawa M (co-first), Hieda M, Taguchi T, Miura N, Kikugawa T, Saika T, *Higashiyama S.
SPOP is essential for DNA-protein crosslink repair in prostate cancer cells: SPOP-dependent removal of topoisomerase 2A from the topoisomerase 2A-DNA cleavage complex.
Mol. Biol. Cell. 31, 478-490 (2020)
PMID: 31967940 - Tezuka-Kagajo M, *Maekawa, M (co-first), *Ogawa A (co-first), Hatta Y, Ishii E, Eguchi M, *Higashiyama S.
Development of human CBF1-targeting single-stranded DNA aptamers with anti-angiogenic activity in vitro.
Nucleic Acid Ther. 30, 365-378 (2020)
PMID: 32881630