Mitsuyasu Kawaguchi
Development of fluorescence probe for proteasomal degradation process imaging and identification of novel proteasome inhibitors
 |
Mitsuyasu Kawaguchi, Ph.DGraduate School of Pharmaceutical Sciences, Nagoya City University, Lab of Bioorganic and Medicinal Chemistry |
|---|
Research summary
Since the development and clinical application of the proteasome inhibitor, bortezomib as an anticancer agent, attention has been focused on the potential efficacy of proteasome inhibitors. However, there have been some cases of resistance to bortezomib, and the development of proteasome inhibitors with new mechanisms of action (MOA) is eagerly awaited. Most of the conventional proteasome inhibitors target the β1, 2, and 5 subunits of the 20S core particle, which exhibit protease activity, and while it is essential to create new target proteins. However, there has been no way to efficiently screening such compounds that regulate novel targets.
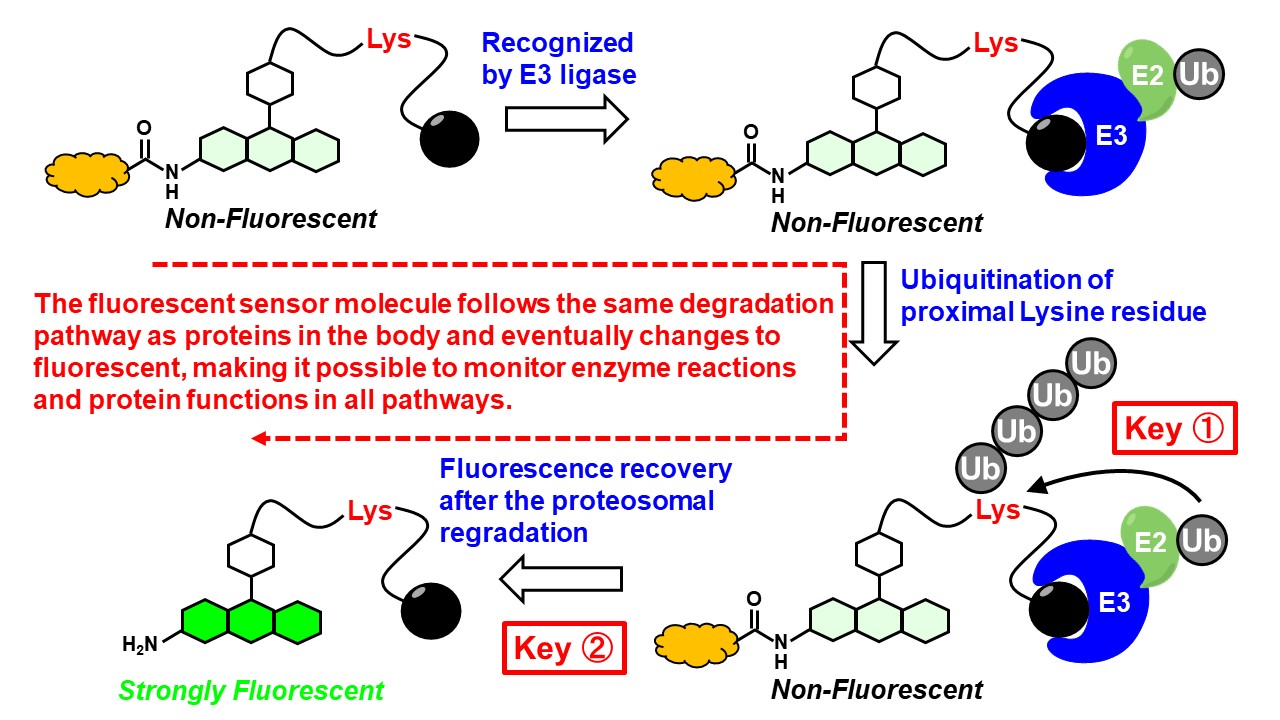
In this study, we devised a completely new detection strategy in which the probe molecule itself is regarded as a substrate of the ubiquitin-proteasome pathway, and becomes fluorescent only when all of the following processes are in progress: i) ubiquitination, ii) incorporation into the 26S proteasome, and iii) proteasomal degradation. Such "fluorescent small molecule probe" can observe the entire proteasomal degradation process in live cells in a single step. We will also apply the developed fluorescent probes to in cellulo phenotypic screening to search for and develop proteasome inhibitors with unprecedented targets and MOA, and demonstrate their efficacy as anticancer drug leads.
In addition, a key question in the development of probes for this research is "does the substrate for ubiquitination have to be a protein? In other words, conventional ubiquitination substrates are usually proteins, however, peptide chains containing lysine residues, lysine residues alone, and even aliphatic amino groups may be sufficient as substrates. By clarifying this question in this study, we may be able to expand the range of application of degradation manipulation technology using PROTACs in chemical biology research, and also present the necessity and opportunity to broaden the view of molecules that are degraded by the ubiquitin-proteasome system in vivo. Furthermore, if we can identify target molecules and regulatory small molecules that have not been identified so far, we will be able to expand the scope of proteasome drug discovery and increase the probability of success in future drug discovery research.
Former Publications
- Kawaguchi M, Ieda N, *Nakagawa H.
Development of peptide-based sirtuin defatty-acylase inhibitors identified by fluorescence probe, SFP3 that can efficiently measure defatty-acylase activity of sirtuin.
J. Med. Chem. 62, 5434-5452 (2019)
PMID: 31117516 - *Kawaguchi M, Han X, Hisada T, Nishikawa S, Kano K, Ieda N, Aoki J, Toyama T, *Nakagawa H.
Development of an ENPP1 fluorescence probe for inhibitor screening, cellular imaging and prognostic assessment of malignant breast cancer.
J. Med. Chem. 62, 9254-9269 (2019)
PMID: 31536342 - Sakamoto S, *Komatsu T, *Watanabe R, Zhang Y, Inoue T, Kawaguchi M, Nakagawa H, Honda K, *Noji H, *Urano Y.
Single enzyme activity-based protein profiling – multiplex single-molecule analysis for counting enzymes for functional identification.
Sci. Adv. 6, eaay0888 (2020)
PMID: 32195342 - *Kawaguchi M, Okabe T, Okudaira S, Hama K, Kano K, Nishimasu H, Nakagawa H, Ishitani R, Kojima H, Nureki O, Aoki J, *Nagano T.
Identification of potent in vivo autotaxin inhibitors that bind to both hydrophobic pocket and channel in the catalytic domain.
J. Med. Chem. 63, 3188-3204 (2020)
PMID: 32134652 - Nakajima Y, *Kawaguchi M, Ieda N, *Nakagawa H.
A Set of Highly Sensitive Sirtuin Fluorescence Probes for Screening Small-Molecular Sirtuin Defatty-Acylase Inhibitors.
ACS Med. Chem. Lett. 12, 617-624 (2021)
PMID: 33859801