Gosuke Hayashi
Ubiquitin chemo-technology powered by chemical protein synthesis and molecular evolution
 |
Gosuke Hayashi, PhDDepartment of Biomolecular Engineering, Graduate School of Engineering, Nagoya University |
|---|
Research summary
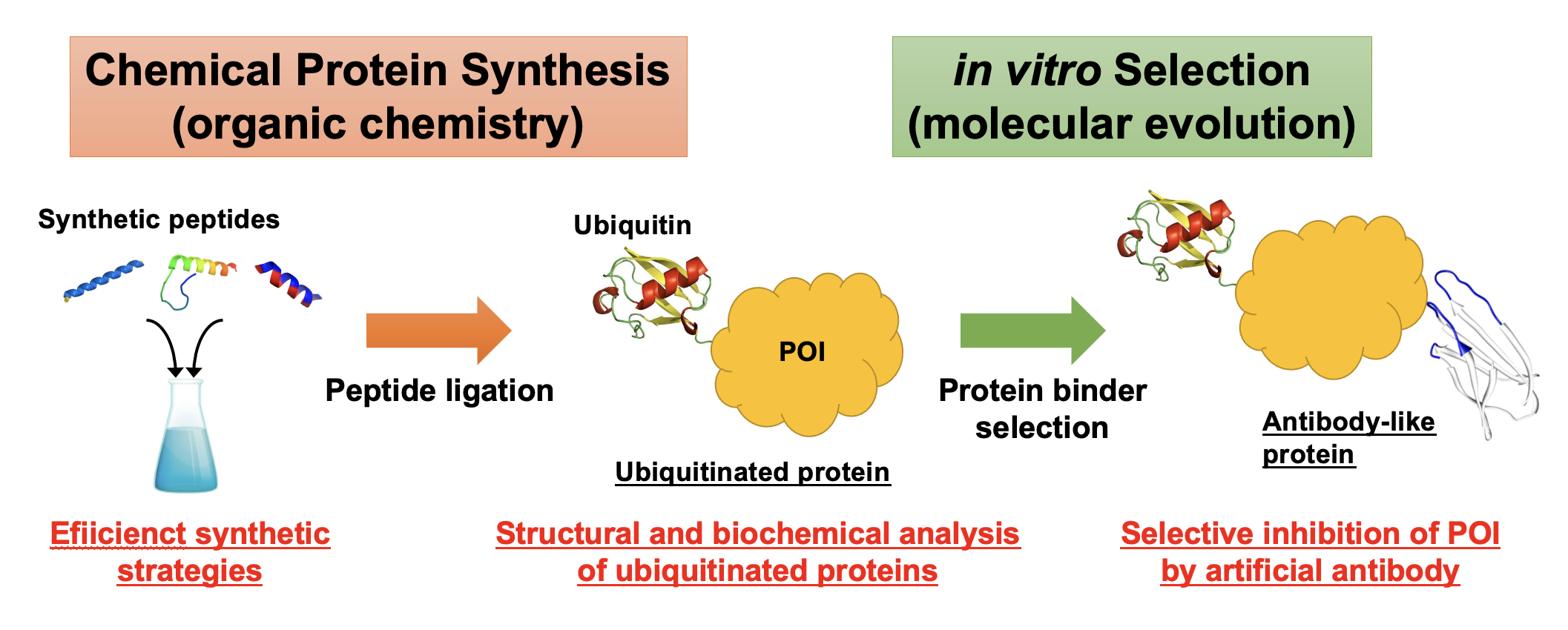
Proteins including ubiquitin are polyamide molecules, in which 20 proteinogenic amino-acids are precisely polymerized, and have been mainly prepared by genetic engineering techniques using ribosomal translation system inside living cells. However, chemical protein synthesis (CPS), in which proteins of interest are prepared through completely chemical process consisting of solid-phase peptide synthesis (SPPS) and following peptide ligation, has been significantly developed in the past two decades. Since various unnatural amino-acids can be site-specifically and multiple incorporated in SPPS procedure, CPS can produce proteins including not only modifications such as posttranslational modifications and/or functional molecules (e.g. fluorescent dyes and crosslinkers) but also unnatural structures such as D-amino acids or N-methyl amino acids.
Our research group has been engaged in developing novel strategies for CPS and applying chemically synthesized proteins into molecular biology researches. For example, we have developed a new one-pot peptide ligation method utilizing organometallic complexes where multiple peptide segments can be continuously assembled without purification steps. In this chemo-ubiquitin project, we combine the CPS techniques with molecular evolution methods such as in vitro selection. Especially, monobody, one of the most popular antibody-like proteins, that specifically and strongly binds to ubiquitinated proteins prepared by CPS or ubiquitin-related proteins, will be developed.
Publications
- Kamo N, *Hayashi G, *Okamoto A.
Silyl-protected propargyl glycine for multiple labeling of peptides by chemoselective silyl-deprotection.
Tetrahedron Lett. 73, 153093 (2021) - Kondo T, Eguchi M, Tsuzuki N, Murata N, Fujino T, Hayashi G, *Murakami H.
Construction of a Highly Diverse mRNA Library for In Vitro Selection of Monobodies.
bio-protocol 11, e4125 (2021)
PMID: 34541043 - Kondo T, Matsuoka K, Umemoto S, Fujino T, Hayashi G, Iwatani Y, *Murakami H.
Monobodies with potent neutralizing activity against SARS-CoV-2 Delta and other variants of concern.
Life Sci. Alliance 5, e202101322 (2022)
PMID: 35256514 - Nakatsu K, *Okamoto A, *Hayashi G, *Murakami H.
Repetitive Thiazolidine Deprotection Using a Thioester-Compatible Aldehyde Scavenger for One-Pot Multiple Peptide Ligation.
Angew. Chem. Int. Ed. 61, e202206240 (2022)
PMID: 35881031 - Akizuki Y, Morita M, Mori Y, Kaiho-Soma A, Dixit S, Endo A, Shimogawa M, Hayashi G, Naito M, Okamoto A, Tanaka K, Saeki Y, *Ohtake, F.
cIAP1-based degraders induce degradation via branched ubiquitin architectures.
Nat. Chem. Biol. 19, 311-322 (2022)
PMID: 36316570 - Hata K, Kobayashi N, Sugimura K, Qin W, Haxholli D, Chiba Y, Yoshimi S, Hayashi G, Onoda H, Ikegami T, Mulholland C, Nishiyama A, Nakanishi M, Leonhardt H, Konuma T, *Arita K.
Structural basis for the unique multifaceted interaction of DPPA3 with the UHRF1 PHD finger.
Nucleic Acid Res. 50, 12527-12542 (2022)
PMID: 36420895
Former Publications
- Kamo N, Hayashi G, *Okamoto A.
Triple Function of 4-Mercaptophenylacetic Acid Promotes One-Pot Multiple Peptide Ligation.
Angew. Chem. Int. Ed. 57, 16533-16537 (2018)
PMID: 30346110 - Yanase M, Nakatsu K, Cardos C J, Konda Y, *Hayashi G, *Okamoto A.
Cysteinylprolyl Imide (CPI) Peptide: A Highly Reactive and Easily Accessible Crypto-thioester for Chemical Protein Synthesis.
Chem. Sci. 10, 5967-5975 (2019)
PMID: 31360403 - Nakatsu K, *Hayashi G, *Okamoto A.
Toolbox for chemically synthesized histone proteins.
Curr. Opin. Chem. Biol. 58, 10-19 (2020)
PMID: 32473259 - Kondo T, Iwatani Y, Matsuoka K, Fujino T, Umemoto S, Yokomaku Y, Ishizaki K, Kito S, Sezaki T, Hayashi G, *Murakami H.
Antibody-like proteins that capture and neutralize SARS-CoV-2.
Sci. Adv. 6, eabd3916 (2020)
PMID: 32948512 - Kamo N, Kujirai T, Kurumizaka H, Murakami H, *Hayashi G, *Okamoto A.
Organoruthenium-Catalyzed Chemical Protein Synthesis to Elucidate the Functions of Epigenetic Modifications on Heterochromatin Factors.
Chem. Sci. 12, 5926-5937 (2021)