Research Group
Research Group
Research group A01: Elucidation of ubiquitin functions by chemo-technologies
Functional regulation of ubiquitin chains using chemo-technologies
Kazuhiro Iwai, MD, PhDDepartment of Molecular and Cellular Physiology, Graduate School of Medicine, Kyoto University |
|
Shiroh Futaki, PhDDivision of Biochemistry, Institute for Chemical Research, Kyoto University |
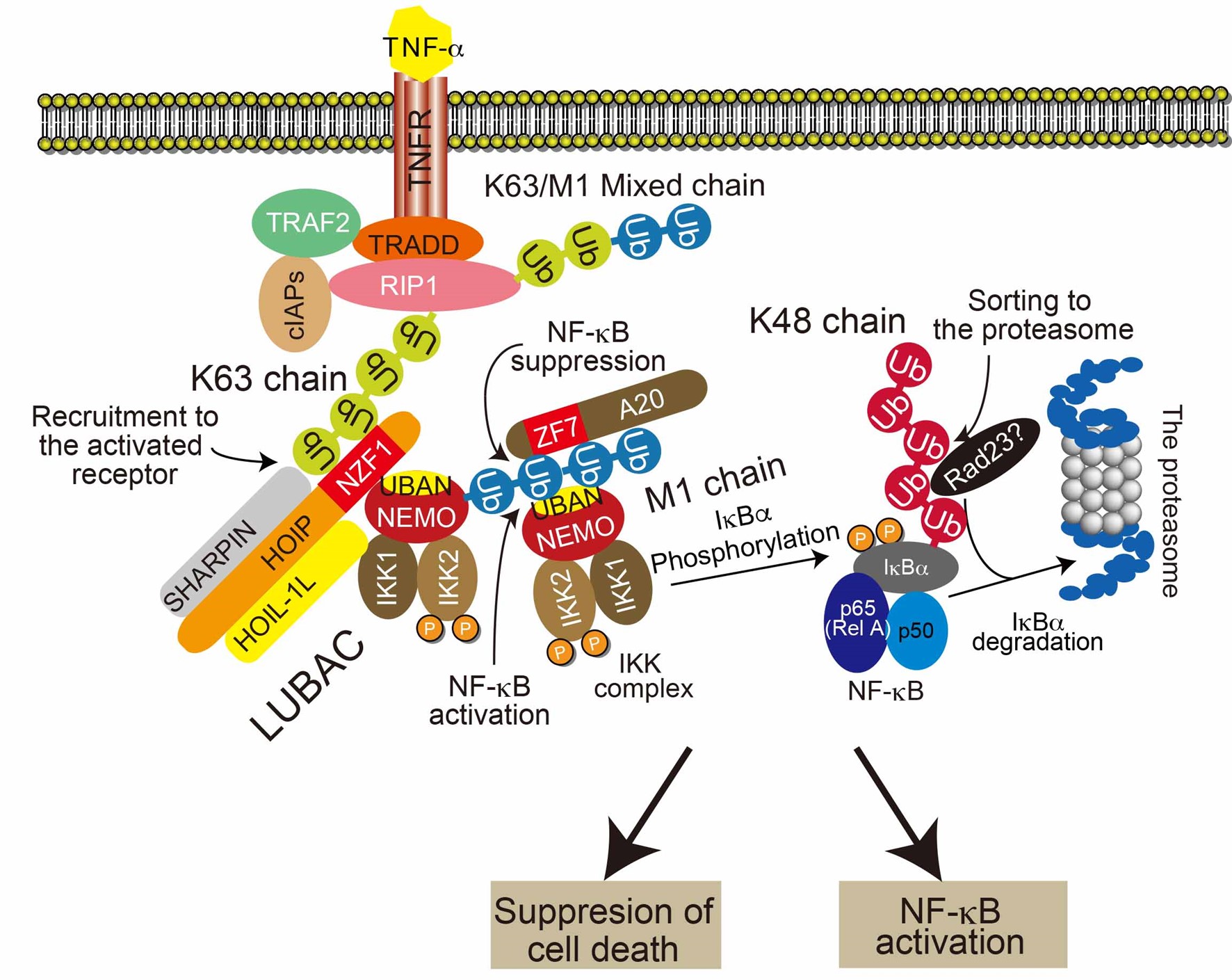
Ubiquitin is conjugated to proteins by a cascade reaction of conjugating enzymes. There are various types ubiquitin modifications and each modification exerts its function by recognized with specific decoder proteins. Cleavage of ubiquitin modifications by deubiquitinases terminates their function. During the research on the linear ubiquitin chain (M1 chain), we realized that the decoder molecules determine the positive or negative signals to some phenomena. For example, recognition of M1 chains by NEMO induces NF-κB activation, whereas A20 attenuates NF-κB by recognizing M1 chains. Therefore, selective inhibition of the binding between ubiquitin modifications and individual decoder molecules is indispensable for detailed functional analyses of ubiquitination. However, such analyses cannot be achieved using currently available methods, and new technological development is awaiting. We found that the short-chain cross-linking peptides (staple peptides) that were generated based on structural information, inhibited tight interaction between two proteins selectively and efficiently. In the current research projects, we will establish a method for selective inhibition for the interaction between ubiquitin modifications and each specific decoder protein using staple peptides in cooperation with the researcher who is well-versed in peptide engineering, and introduce a powerful new analytical tool for ubiquitin research. Various ubiquitin modifications are involved in the TNF-α signaling pathway. Using the TNF-α signal system as one of the model systems, we will promote our projects to establish a new chemo-technology specific for ubiquitin research and understand ubiquitin biology using the new chemical tools.
Publications
- *MacDuff DA, Baldridge MT, Qaqish AM, Nice TJ, Darbandi AD, Hartley VL, Peterson ST, Miner JJ, Iwai K, *Virgin HW.
HOIL1 is essential for the induction of type I and III interferons by MDA5 and regulates persistent murine norovirus infection.
J. Virol. 92, e01368-18 (2018)
PMID: 30209176 - Wu M, Chang Y, Hu H, Mu R, Zhang Y, Qin X, Duan X, Li W, Tu H, Zhang W, Wang G, Han Q, Li A, Zhou T, Iwai K, Zhang X *Li H.
LUBAC controls chromosome alignment by targeting CENP-E to attached kinetochores.
Nat. Commun. 10, 273 (2019)
PMID: 30655516 - Ishii N, Walinda E, Iwakawa N, Morimoto D, Iwai K, Sugase K, *Shirakawa M.
NMR resonance assignments of the NZF domain of mouse HOIL-1L free and bound to linear di-ubiquitin.
Biomol. NMR Assign. 13, 149-153 (2019)
PMID: 30569274 - Sasaki K, Himeno A, Nakagawa T, Sasaki Y, Kiyonari H, *Iwai K.
Modulation of autoimmune pathogenesis by T cell-triggered inflammatory cell death.
Nat. Commun. 10, 3878 (2019)
PMID: 31462647 - Kawaguchi Y, Ise S, Azuma Y, Takeuchi T, Kawano K, Le TK, Ohkanda J, *Futaki S. Dipicolylamine/metal complexes that promote direct cell-membrane penetration of octaarginine.
Bioconjug. Chem. 30, 454-460 (2019)
PMID: 30428256 - Brazee PL, Morales-Nebreda L, Magnani ND, Garcia JG, Misharin AV, Ridge KM, Budinger GRS, Iwai K, Dada LA, *Sznajder JI.
Linear ubiquitin assembly complex regulates lung epithelial-driven responses during influenza infection.
J. Clin. Invest. 130, 1301-1314 (2020)
PMID: 31714898 - Arafiles JVV, Hirose H, Akishiba M, Tsuji S, Imanishi M, *Futaki S.
Stimulating macropinocytosis for intracellular nucleic acid and protein delivery: a combined strategy with membrane-lytic peptides to facilitate endosomal escape.
Bioconjug. Chem. 31, 547-553 (2020)
PMID: 32017537 - Tamemoto N, Akishiba M, Sakamoto K, Kawano K, Noguchi H, *Futaki S.
Rational Design Principles of Attenuated Cationic Lytic Peptides for Intracellular Delivery of Biomacromolecules.
Mol. Pharm. 17, 2175-2185 (2020)
PMID: 32352304 - Sakamoto K, Akishiba M, Iwata T, Murata K, Mizuno S, Kawano K, Imanishi M, Sugiyama F, *Futaki S.
Optimizing Charge Switching in Membrane Lytic Peptides for Endosomal Release of Biomacromolecules.
Angew. Chem. Int. Ed. Engl. 59, 19990-19998 (2020)
PMID: 32557993 - Fuseya Y, Fujita H, Kim M, Ohtake F, Nishide A, Sasaki K, Saeki Y, Tanaka K, Takahashi R, Iwai K.
The HOIL-1L ligase modulates immune signaling and cell death via mono-ubiquitination of LUBAC.
Nat. Cell Biol. 22, 663-673 (2020)
PMID: 32393887 - Jo T, *Nishikori M, Kogure Y, Arima H, Sasaki K, Sasaki Y, Nakagawa T, Iwai F, Momose S, Shiraishi A, Kiyonari H, Kagaya N, Onuki T, Shin-ya K, Yoshida M, Kataoka K, Ogawa S, *Iwai K, Takaori-Kondo A.
LUBAC accelerates B-cell lymphomagenesis by conferring B cells resistance to genotoxic stress.
Blood. 136, 684–697 (2020)
PMID: 32325488 - Arafiles JVV, Hirose H, Hirai Y, Kuriyama M, Sakyiamah MM, Nomura W, Sonomura K, Imanishi M, Otaka A, Tamamura H, *Futaki S.
Discovery of a macropinocytosis-inducing peptide potentiated by medium-mediated intramolecular disulfide formation.
Angew. Chem. Int. Ed. Engl. 60,11928-11936 (2021)
PMID: 33629482 - Morimoto D, Walinda E, Takashima S, Nishizawa M, Iwai K, Shirakawa M, *Sugase K.
Structural dynamic heterogeneity opolyubiquitin subunits affects phosphorylation susceptibility.
Biochemistry 60, 573-583 (2021)
PMID: 33616406 - Yoshida Y, Asahina M, Murakami A, Kawawaki J, Yoshida M, Fujinawa R, Iwai K, Tozawa R, Matsuda N, Tanaka K and Suzuki T.
Loss of peptide:N-glycanase causes proteasome dysfunction mediated by a sugar-recognizing ubiquitin ligase.
Proc. Natl. Acad. Sci. USA 118, e2102902118 (2021)
PMID: 34215698 - Walinda E, Morimoto D, Sorada T, Iwai K, Sugase K.
Expression, solubility monitoring, and purification of the co-folded LUBAC LTM domain by structure-guided tandem folding in autoinducing cultures.
Protein Expr. Purif. 187,105953 (2021)
PMID: 34390872 - Iwata T, Hirose H, Sakamoto K, Hirai Y, Arafiles JVV, Akishiba M, Imanishi M, *Futaki S.
Liquid Droplet Formation and Facile Cytosolic Translocation of IgG in the Presence of Attenuated Cationic Amphiphilic Lytic Peptides.
Angew. Chem. Int. Ed. Engl. 60, 19804-19812 (2021)
PMID: 34114295 - Nitschke S, Sullivan MA, Mitra S, Marchioni CR, Lee JPY, Smith BH, Ahonen S, Wu J, Chown EE, Wang P, Petković S, Zhao X, DiGiovanni LF, Perri AM, Israelian L, Grossman TR, Kordasiewicz H, Vilaplana F, Iwai K, Nitschke F, Minassian BA.
Glycogen synthase downregulation rescues the amylopectinosis of murine RBCK1 deficiency.
Brain, awac017 (2022)
PMID: 35084461 - Shinkawa Y, Imami K, Fuseya Y, Sasaki K, Ohmura K, Ishihama Y, Morinobu A, Iwai K.
ABIN1 is a signal-induced autophagy receptor that attenuates NF-kB activation by recognizing linear ubiquitin chains.
FEBS Lett. 596, 1147-1164 (2022)
PMID: 35213742 - Wood MJ, Marshall JN, Hartley VL, Liu TC, Iwai K, Stappenbeck TS and MacDuff DA. HOIL1 regulates group 2 innate lymphoid cell numbers and type 2 inflammation in the small intestine.
Mucosal Immunol. 15, 642-655 (2022)
PMID: 35534698 - Nakagawa Y, Arafiles JVV, Kawaguchi Y, Nakase I, Hirose H, Futaki S.
Stearylated Macropinocytosis-Inducing Peptides Facilitating the Cellular Uptake of Small Extracellular Vesicles.
Bioconjug. Chem. 33, 869-880 (2022)
PMID: 35506582 - Okano S, Kawaguchi Y, Kawano K, Hirose H, Imanishi M, Futaki S.
Split luciferase-based estimation of cytosolic cargo concentration delivered intracellularly via attenuated cationic amphiphilic lytic peptides.
Bioorg .Med. Chem. Lett. 72, 128875 (2022)
PMID: 35798239 - Hirose H, Hirai Y, Sasaki M, Sawa H, Futaki S.
Quantitative Analysis of Extracellular Vesicle Uptake and Fusion with Recipient Cells.
Bioconjug. Chem. 33, 1852-1859 (2022)
PMID: 36194183 - Jimbo K, Hattori A, Koide S, Ito T, Sasaki K, Iwai K, Nannya Y, Iwama A, Tojo A, and Konuma T.
Genetic deletion and pharmacologic inhibition of E3 ubiquitin ligase HOIP impairs the propagation of myeloid leukemia.
Leukemia 37, 122-133 (2023)
PMID: 36352193 - Hiragi K, Nishide A, Takagi K, Iwai K, Kim M, and Mizushima T.
Structural insight into the recognition of the linear ubiquitin assembly complex by Shigella E3 ligase IpaH1.4/2.5.
J. Biochem. (2023) Jan 4:mvac109.
PMID: 36610722 - Tokunaga F, Sakata S-I, Saeki Y, Satomi Y, Kirisako T, Kamei K, Nakagawa T, Kato M, Murata S, Yamaoka S, Yamamoto M, Akira S, Takao T, Tanaka K, Iwai K.
Involvement of linear polyubiquitination of NEMO in NF-kB activation.
Nat. Cell Biol. 11, 123-132 (2009)
PMID :19136968 - Tokunaga F, Nakagawa T, Nakahara M, Saeki Y, Taniguchi M, Sataka S-I, Tanaka K, Nakano H, Iwai K.
SHARPIN is a component of the NF-kB activating linear ubiquitin chain assembly complex.
Nature 471, 633-636 (2011)
PMID: 21455180 - Fujita H, Rahighi S, Akita M, Kato R, Sasaki Y, Wakatsuki S, Iwai K.
Mechanism underlying IKK activation mediated by the linear ubiquitin chain assembly complex (LUBAC).
Mol. Cell. Biol. 34, 1322-1335 (2014)
PMID: 24469399 - Iwai K, Fujita H, Sasaki Y.
Linear ubiquitin chains: NF-kB signalling, cell death, and beyond.
Nat. Rev. Mol. Cell Biol. 15, 503-508 (2014)
PMID: 25027653 - Yang Y, Schmitz R, Mitala J, Whiting A, Xiao W, Ceribelli M, Wright GW, Zhao H, Yang Y, Xu W, Rosenwald A, Ott G, Gascoyne RD, Connors JM, Rimsza LM, Campo E, Jaffe ES, Delabie J, Smeland EB, Braziel RM, Tubbs RR, Cook JR, Weisenburger DD, Chan WC, Wiestner A, Kruhlak MJ, Iwai K, Bernal F, Staudt LM.
Essential role of the linear ubiquitin chain assembly complex in lymphoma revealed by rare germline polymorphisms.
Cancer Discov. 4, 480-493 (2014)
PMID: 24491438 - Shimizu S, Fujita H, Sasaki Y, Tsuruyama T, Fukuda K, Iwai K.
Differential Involvement of the Npl4 Zinc Finger Domains of SHARPIN and HOIL-1L in Linear Ubiquitin Chain Assembly Complex-Mediated Cell Death Protection.
Mol. Cell. Biol. 36, 1569-1583 (2016)
PMID: 26976635 - Nakase I, Noguchi K, Aoki A, Takatani-Nakase T, Fujii I, Futaki S.
Arginine-rich cell-penetrating peptide-modified extracellular vesicles for active macropinocytosis induction and efficient intracellular delivery.
Sci. Rep. 7, 1991 (2017)
PMID: 28512335 - Akishiba M, Takeuchi T, Kawaguchi Y, Sakamoto K, Yu HH, Nakase I, Takatani-Nakase T, Madani F, Gräslund A, Futaki S.
Cytosolic antibody delivery by lipid-sensitive endosomolytic peptide.
Nat. Chem. 9, 751-761 (2017)
PMID: 28754944 - Futaki S, Nakase I.
Cell-surface interactions on arginine-rich cell-penetrating peptides allow for multiplex modes of internalization.
Acc. Chem. Res. 50, 2449-2456 (2017)
PMID: 28910080 - Tamiya H, Kim H, Klymenko O, Kim H, Feng Y, Zhang T, Han JY, Murao A, Snipas SJ, Jilaveanu L, Brown K, Kluger H, Zhang H, Iwai K, Ronai ZA.
SHARPIN-mediated regulation of protein arginine methyltransferase 5 controls melanoma growth.
J. Clin. Invest. 128, 517-530 (2018)
PMID: 29227283 - Sakagami K, Masuda T, Kawano K, Futaki S.
Importance of net hydrophobicity in the cellular uptake of all-hydrocarbon stapled peptides.
Mol. Pharm. 15, 1332-1340 (2018)
PMID: 29420899 - Fujita H, Tokunaga A, Shimizu S, Whiting AL, Aguilar-Alonso F, Takagi K, Walinda E, Sasaki Y, Shimokawa T, Mizushima T, Ohki I, Ariyoshi M, Tochio H, Bernal F, Shirakawa M, Iwai K.
Cooperative domain formation by homologous motifs in HOIL-1L and SHARPIN plays crucial roles in LUBAC stabilization.
Cell Rep. 23, 1192-1204 (2018)
PMID: 29694895